OUR TECHNOLOGIES
We evaluate enzymatic activity at single protein level.
We can live well when our "cells" are working properly. Within cells are “proteins”, which consist of several types including enzymes. Enzymes play various important roles in “cells” function. For example, if the enzyme that acts as scissors cuts more substrates than usual, or, conversely, stops cutting at all, the cells will not work well. Often times, this change in enzyme activity is known to cause or worsen disease. Thus, we can accurately detect disease by evaluating the enzyme performance or activity at each enzyme level.
Single-molecule Enzyme Activity-Based Liquid Biopsy
STEP1)
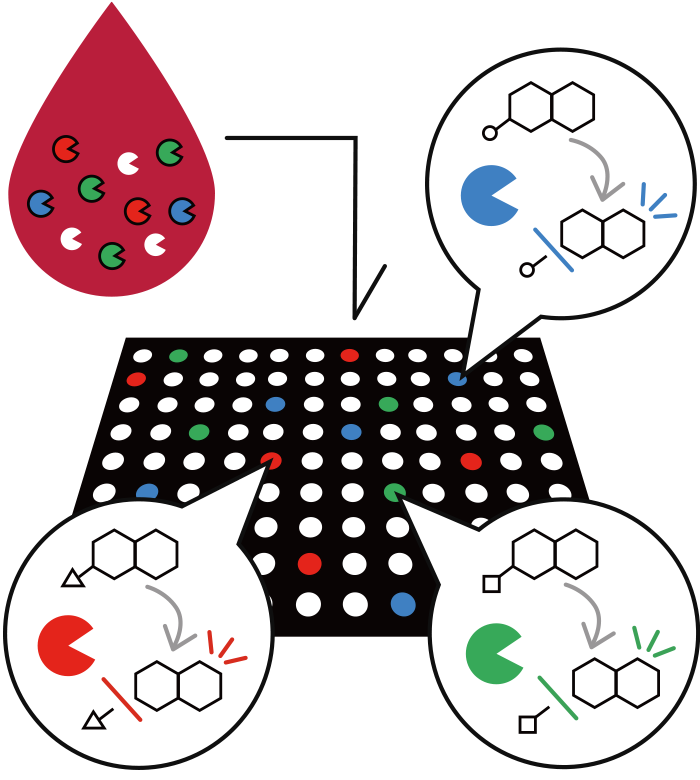
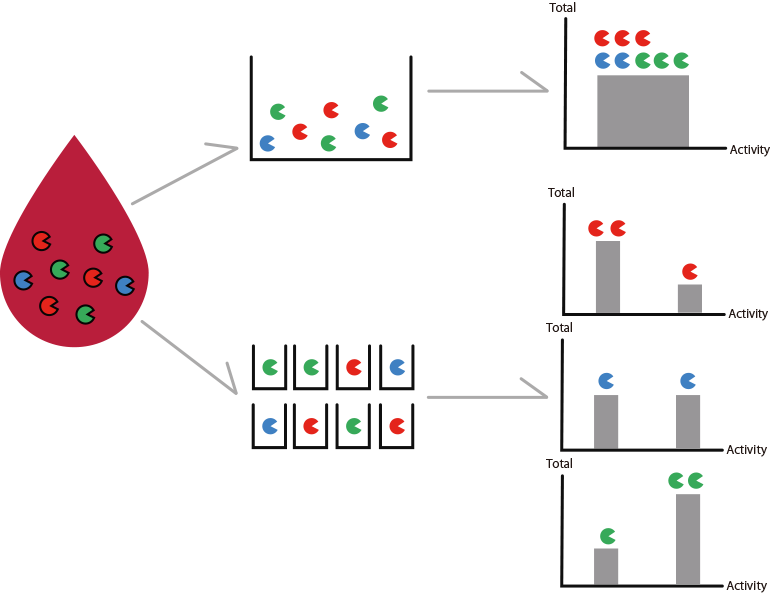
Diluted blood with multi-colored fluorescent enzyme substrates are added to a microdevice with numerous wells with a diameter of 3 µm. Probability wise, each well will include one or zero enzyme. The enzymes metabolize the fluorescent enzyme substrates, producing fluorescent material in the wells. Fluorescence microscopy provides fluorescent signals with an intensity corresponding to the activity of each enzyme.
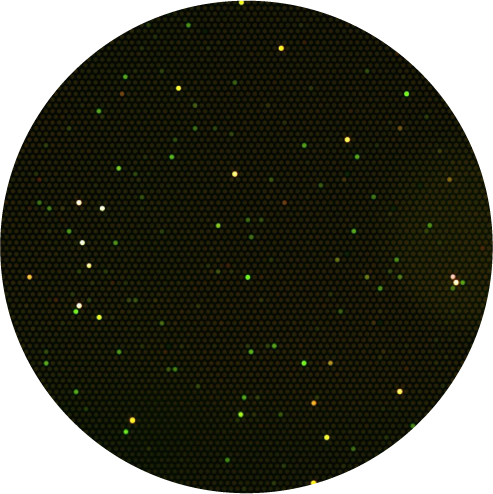
STEP2)
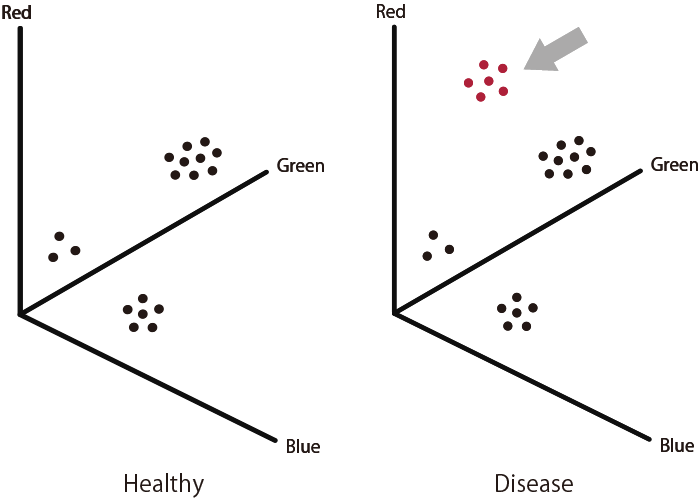
Diseases such as cancers are diagnosed based on difference in enzyme activity profiles, such as number of enzymes (based on number of glowing wells) and enzyme activity (fluorescence intensity of each color).
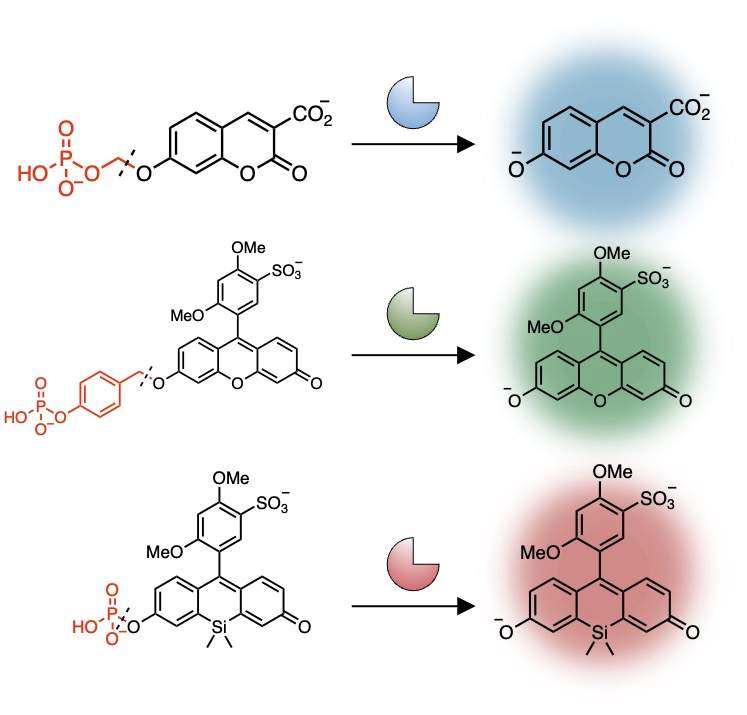
Fluorescent enzyme substrates (fluorescent probes)
We designed and developed our own fluorescent enzyme substrates that fluoresce when metabolized by certain enzyme groups.
To date, more than 100 fluorescent enzyme substrates have been developed, including ALP (Alkaline phosphatases), ENPP (Ectonucleotide pyrophosphatases/phosphodiesterases), MMP (Matrix metalloproteinases), DPP (Dipeptidyl peptidases), Aminopeptidases, etc.
Enzyme subtypes can be distinguished by the differences in the enzyme reactivity to the fluorescent enzyme substrate of each color.
THREE ADVANTAGES
1. High predictive accuracy
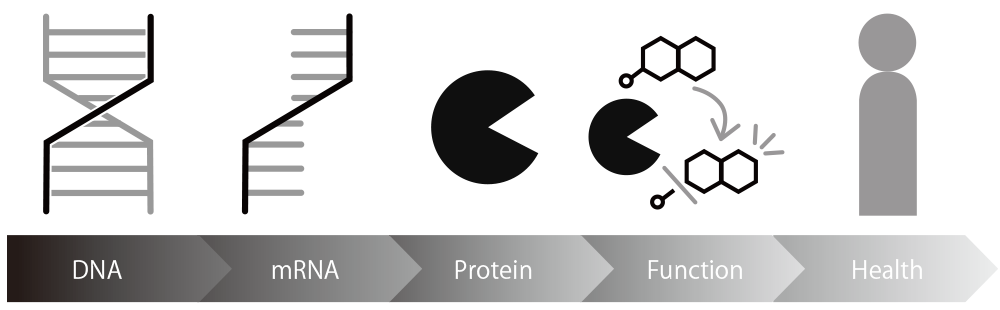
Existing diagnostics mostly detect proteins which include DNA, mRNA, and enzymes, but detecting the change levels of such proteins may not accurately capture changes in disease state.
Our technology utilizes "enzyme activity" which is directly related to cellular function and is thought to more accurately reflect the disease state as an indicator.
Diagnoses with high predictive accuracy is made possible by capturing the intensity of activity for each molecule of enzyme.
2. High detection sensitivity
For example, in the conventional ELISA method, 10 million enzymes are required for detection.
Our technology enables enzymes to be detected at single enzyme level with a microdevice, thus allowing for earlier diagnosis.
3. Detectable with less than a drop of blood
Since 1 µL of blood is sufficient for diagnosis, evaluation is possible with a very small amount of sample.

RESEARCH ACHIEVEMENTS
This is the result of our CTO Dr. Sakamoto's graduate school work. Utilizing the innovative Single-molecule Enzyme Activity-Based Enzyme Profiling (SEAP) technology, which combines fluorescent enzyme substrates and microdevices, we have successfully demonstrated the capability to detect and differentiate the activity of various phosphate hydrolysis enzymes in blood at the single-molecule level. Moreover, our research revealed a significant increase in the size of ENPP3 activity clusters in the blood of pancreatic cancer patients compared to healthy individuals. This discovery highlights the potential of our technique in revolutionizing disease diagnosis.
Our partner at the Universitry of Tokyo has created an innovative fluorescent enzyme substrate capable of detecting M3 metalloprotease activity. They observed an elevation in enzyme activity within the cancerous tissues and blood samples of individuals with colorectal cancer. This finding underscores the potential utility of Single-molecule Enzyme Activity-based Liquid Biopsy in the early detection of colorectal cancer.
This is the research achievement towards founding Cosomil under the JST START grant project. We have developed an innovative method to analyze the activity of various proteolytic enzymes in blood at the single-molecule level. Through this technique, we've identified abnormal enzyme activities such as DPP4 and CD13 in the blood of Stage I-II pancreatic cancer patients. Leveraging these insights, we are advancing our pioneering Single-molecule Enzyme Activity-based Liquid Biopsy technology for the early detection of pancreatic cancer.
This work is the result of a collaborative research project with the Graduate School of Pharmaceutical Sciences at the University of Tokyo. By using an NAD(P)H-responsive fluorescent probe optimized for microdevices, we developed a new methodology to analyze enzyme activity of oxidoreductases in body fluids at the single-molecule level. These findings enables us to discover disease biomarkers in a new category of enzymes in addition to the proteases.
